SYSTCM offers two methods to browse data, including 550 Traditional Chinese Medicines, 10936 components, 40 pharmacological effects, 18 efficacies and 18 TCM properties. After clicking, it will jump to the corresponding page.
(1) Navigation bar at the top of any page

(2) In "Picture" on the home page.
On every page, users can search the different types of data provided by SYSTCM. Firstly, users should select the type that they want to retrieve. Secondly, please enter terms into the input box, and click the "Search" button or press "ENTER" to jump to the search results page.

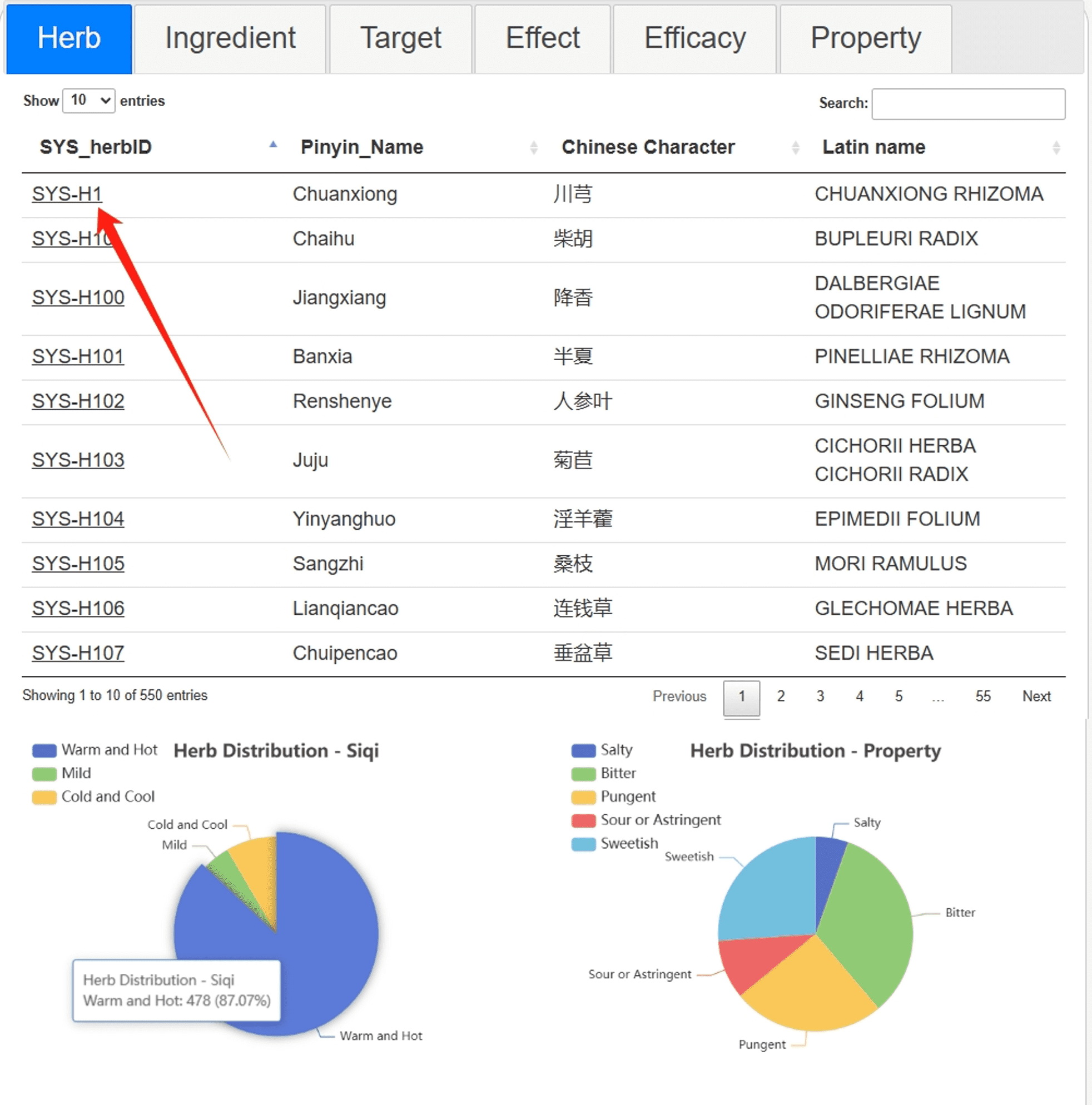
On the Browse page, data from six categories of SYSTCM is displayed. A pie chart below provides a simple statistical classification of the data. By clicking on an entry number of interest, you can access the detailed information page.
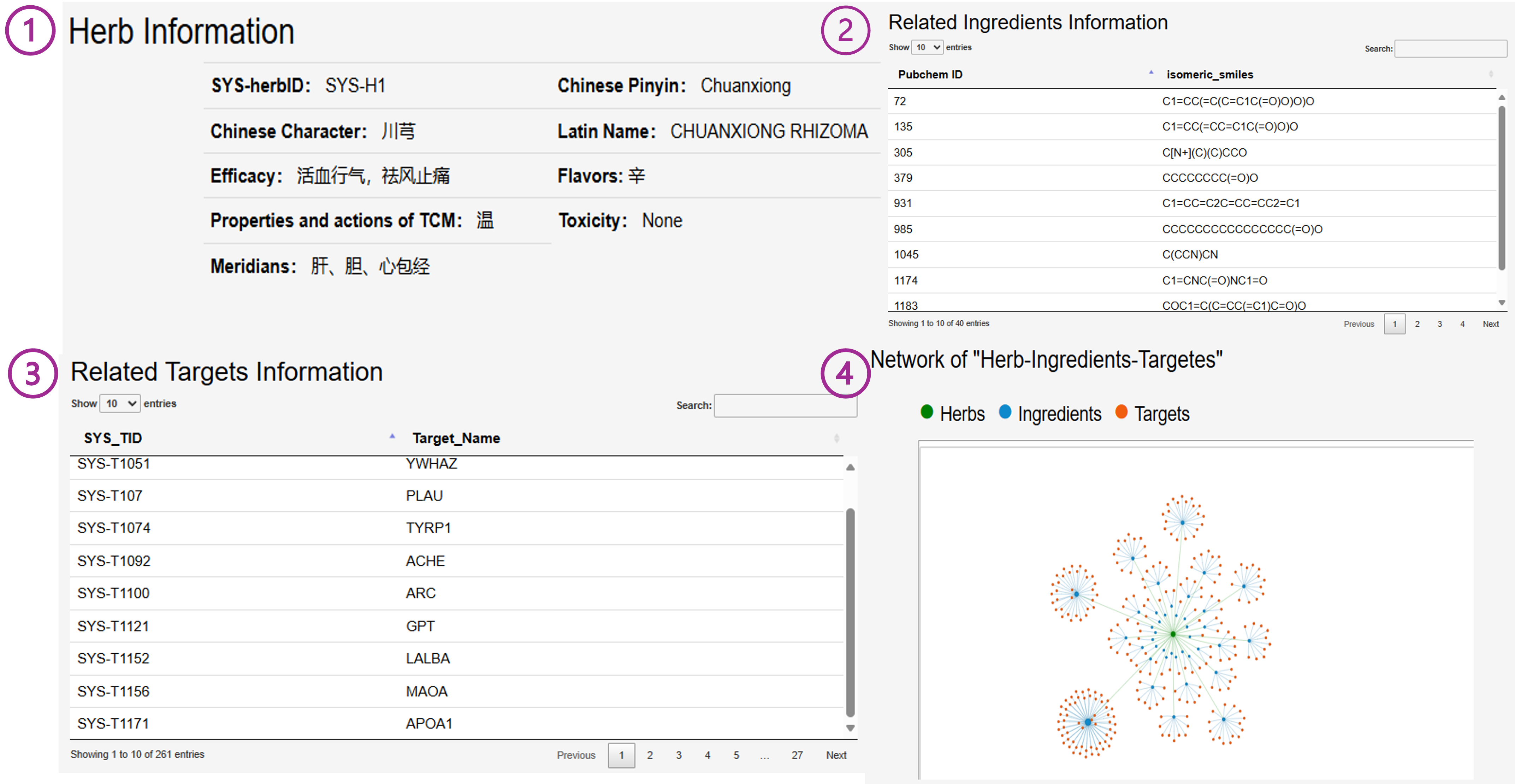
On the Details page, the detailed information of the selected entry and related entries is displayed, along with a network diagram for visualization. For example, in the case of Herb, the page presents its basic information, such as ID, the Four Qi, Five Flavors, and Meridians it belongs to. Additionally, it displays the components contained in the herb, their target proteins, and the Herb-Ingredients-Targets network diagram.
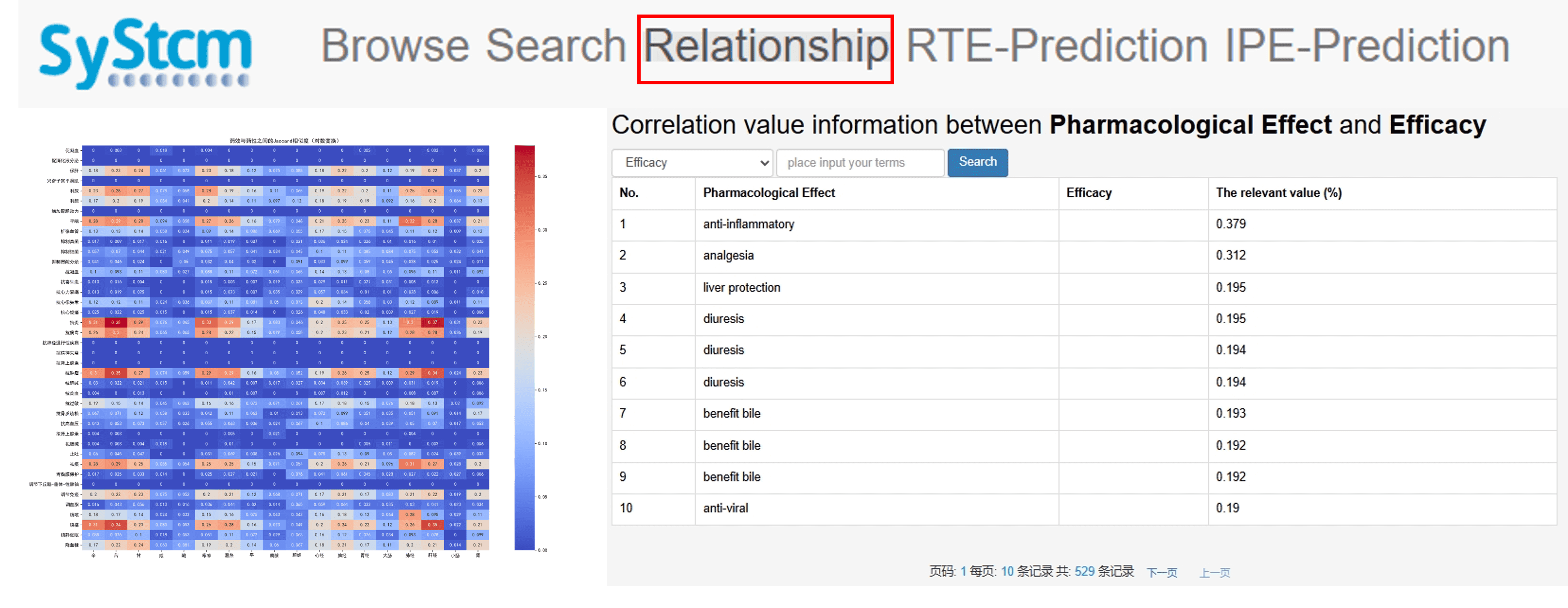
In the Relationship section, we analyze the associations between the number of literature records retrieved from CNKI for specific traditional Chinese medicines and their efficacy and medicinal properties as recorded in the Chinese Pharmacopoeia. By calculating the Jaccard similarity between these datasets, we determine the pairwise association values among 40 pharmacological effects, 18 therapeutic efficacies, and 18 medicinal properties.
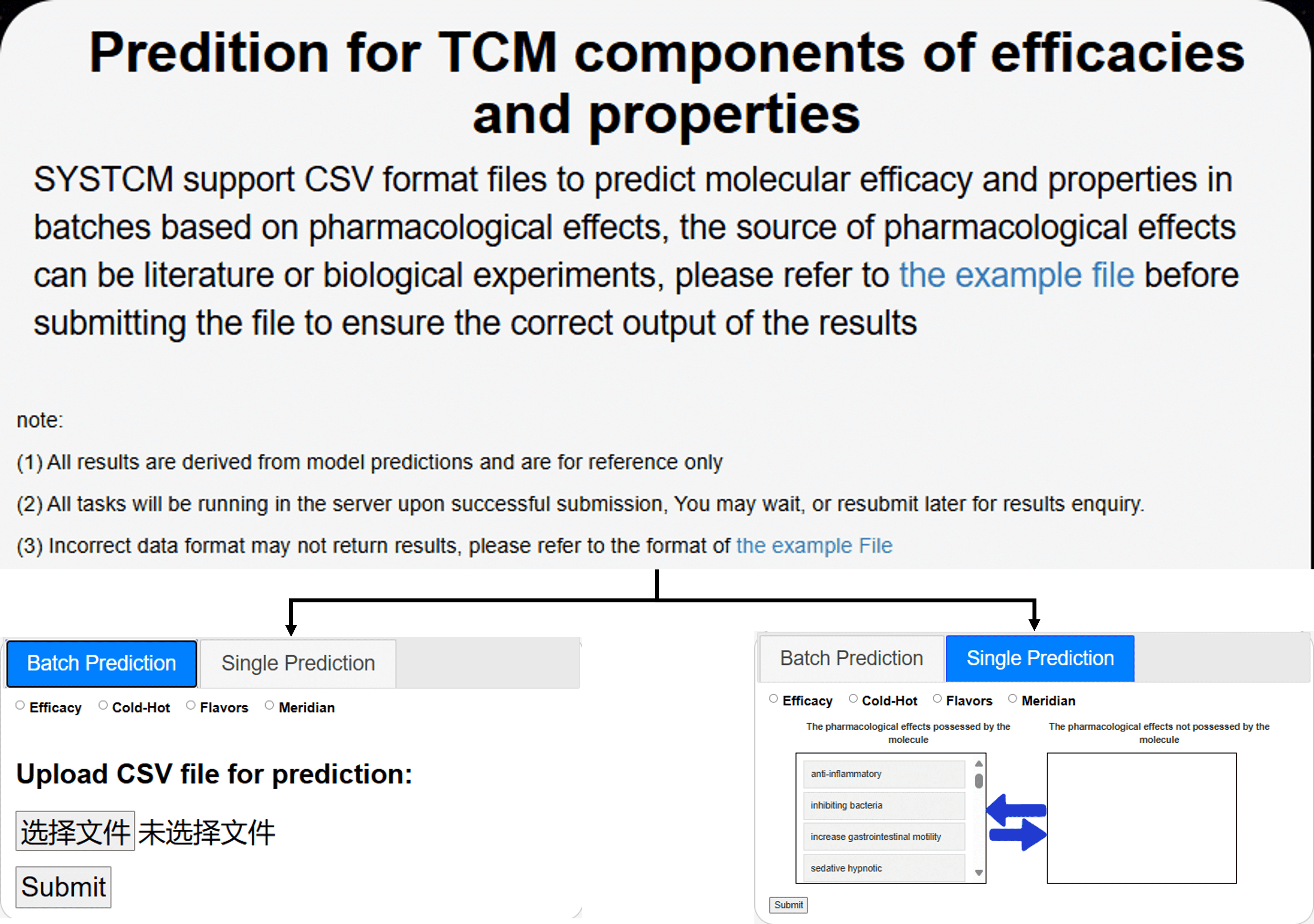
5.1 Prediction for TCM components of efficacies and properties
SYSTCM supports both batch and single prediction of the efficacy and properties of components, constituent, or herb.
For batch prediction, users can click on "the example file"> to view a sample file. By following the format, they can specify whether certain samples possess specific pharmacological effects. Users can then upload a CSV file, select the desired prediction targets (efficacy, cold-hot properties, five flavors and so on), and submit it to SYSTCM for batch processing.
For single prediction, users can directly select the prediction target, specify whether the sample has or does not have certain pharmacological effects in the text box below, and submit it to SYSTCM for prediction.
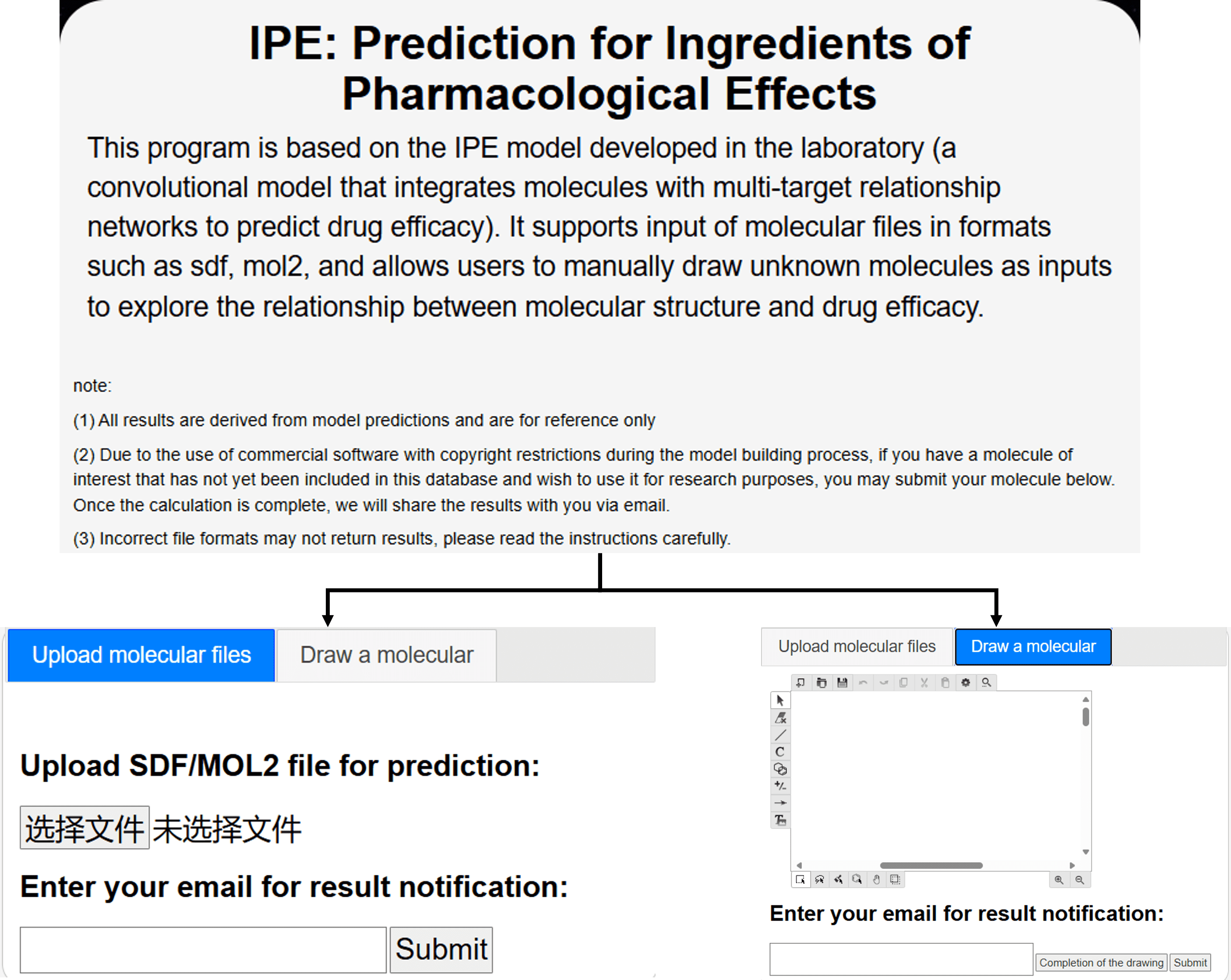
5.2 Prediction for Ingredients of Pharmacological Effects
The effects prediction module can be accessed by clicking on the IPE-Prediction module in the navigation bar.
module is developed based on the convolutional neural network (CNN). It takes molecular structure data as input and predicts the presence or absence of 40 pharmacological effects for each sample. The module supports both batch predictions and single predictions. Regardless of the prediction type, SYSTCM requires users to upload all samples in a single file (e.g., SDF or MOL2 format), which will be processed by the system in the background.
For newly discovered or modified molecular structures, SYSTCM integrates the Kekulé molecular editor, enabling users to draw, modify, save, and export molecular structures. This allows users to upload customized molecular samples for efficacy prediction. Additionally, when submitting molecular data, users are required to provide an email address. The laboratory staff will periodically process the submissions and send the results to users via email.
